How to Perform a Shapiro-Wilk Normality Test in R (2 Examples)
This article shows how to perform a Shapiro-Wilk normality test in the R programming language.
Creation of Example Data
data(iris) # Iris data set as example data head(iris) # Sepal.Length Sepal.Width Petal.Length Petal.Width Species # 1 5.1 3.5 1.4 0.2 setosa # 2 4.9 3.0 1.4 0.2 setosa # 3 4.7 3.2 1.3 0.2 setosa # 4 4.6 3.1 1.5 0.2 setosa # 5 5.0 3.6 1.4 0.2 setosa # 6 5.4 3.9 1.7 0.4 setosa |
data(iris) # Iris data set as example data head(iris) # Sepal.Length Sepal.Width Petal.Length Petal.Width Species # 1 5.1 3.5 1.4 0.2 setosa # 2 4.9 3.0 1.4 0.2 setosa # 3 4.7 3.2 1.3 0.2 setosa # 4 4.6 3.1 1.5 0.2 setosa # 5 5.0 3.6 1.4 0.2 setosa # 6 5.4 3.9 1.7 0.4 setosa
Example 1: Using shapiro.test Function to Conduct Shapiro-Wilk Normality Test
shapiro.test(iris$Sepal.Length) # Applying shapiro.test function # Shapiro-Wilk normality test # # data: iris$Sepal.Length # W = 0.97609, p-value = 0.01018 |
shapiro.test(iris$Sepal.Length) # Applying shapiro.test function # Shapiro-Wilk normality test # # data: iris$Sepal.Length # W = 0.97609, p-value = 0.01018
The Shapiro-Wilk test shows a significant result considering a 0.05 p-value level, i.e. the variable Sepal.Length is not normally distributed.
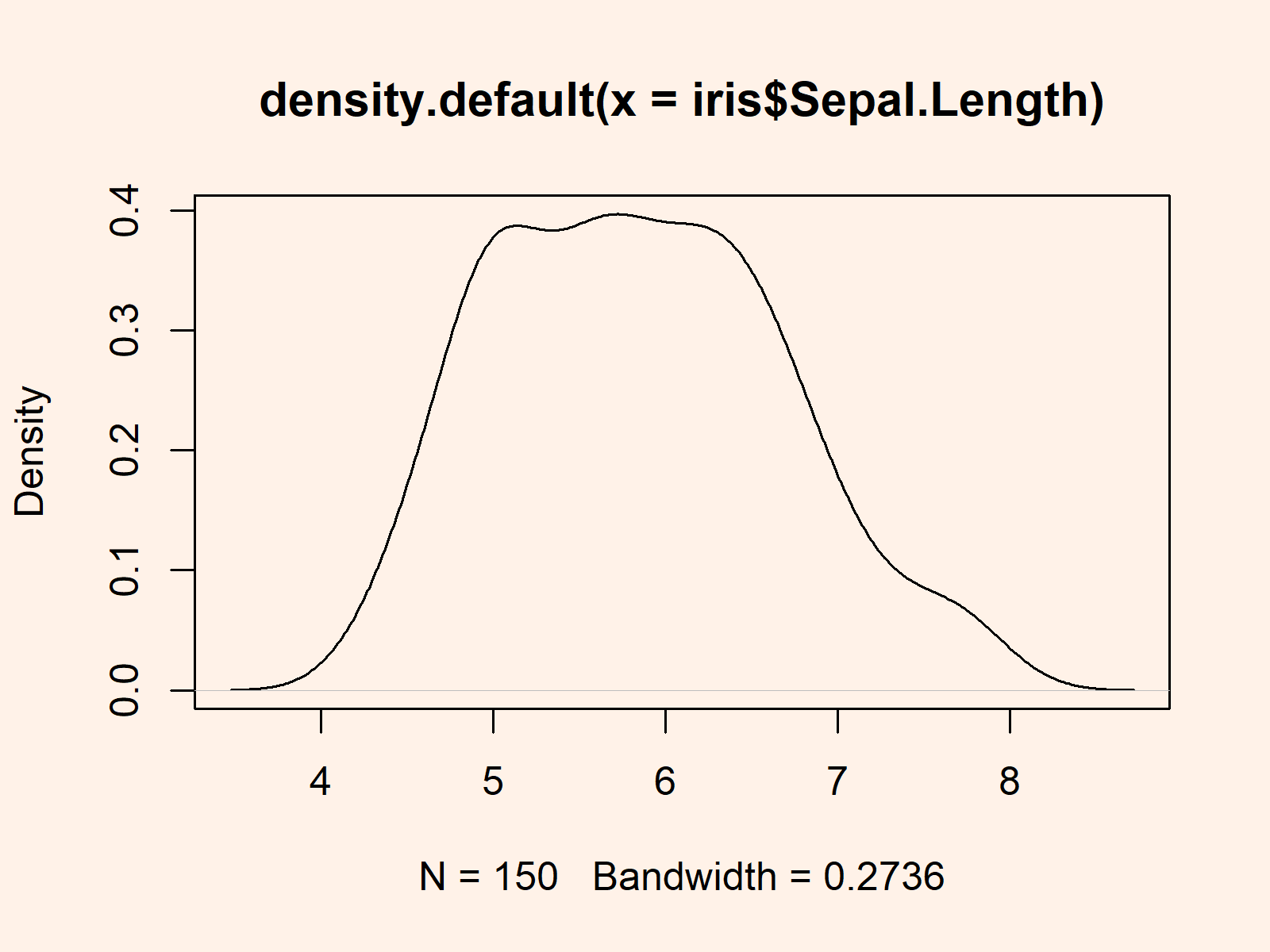
Example 2: Drawing Density Plot of Data
plot(density(iris$Sepal.Length)) # Non-normal density |
plot(density(iris$Sepal.Length)) # Non-normal density